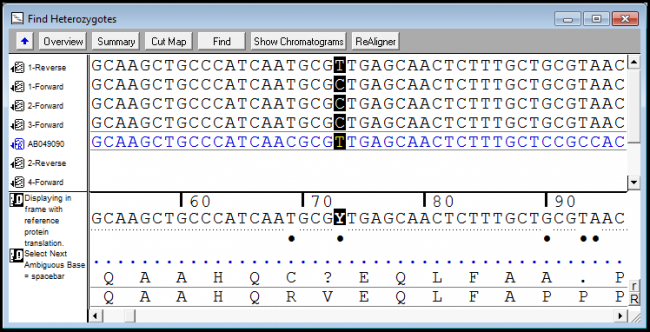
The excess of heterozygous variants should be. DNA sequencing encompasses biochemical methods for determining the order of the nucleotide bases, in a DNA oligonucleotide.DNA sequence determines the patterns that make up genetic traits and in some cases behaviors. Which will bring you to square one: look at your Sanger chromatograms. Our results show that rare causative mutations in known ARNSHL genes can be reliably identified via WES. You are still out of luck if instead of clear homozygote different from reference you will get "N" base in your fasta/fastq file, but not just one but multiple reads.
#DNA SEQUENCING SEQUENCHER HETEROZYGOUS MUTATIONS SOFTWARE#
reference sequences using Sequencher software (Gene Codes Corporation, Ann Arbor. Staden package listed above, then you have to have some genome sequence from your species, even if it is a toy "genome" in a form of contigs of interest, convert your Sanger files to FASTQ (i.e by using BioPython: ), or to FASTA, then map it to your toy genome and load into IGV. We have identified a novel heterozygous mutation in XPD (c.1906C>T p. In case you have a lot of Sanger sequences plus FASTQ from NGS, and simply refuse to use i.e. Methods and results: The direct sequencing method was improved by devising primers for amplifying the LPL gene and. Sometimes sequencing run is longer than your insert or gets killed for other reasons, and you get random noise base calls. Objective: The purpose of this study was to develop an improved method of direct DNA sequencing, which makes it possible to identify heterozygous mutations of the lipoprotein lipase (LPL) gene in order to understand the underlying genetic disorder of type IV hyperlipoproteinemia. If you can not see the chromatogram, and use simple fasta you can not make any calls as what is going on regarding SNPs/indels. There are specific tools for viewing Sanger chromatograms and it makes no sense try to load these chromatogram files into programs which simply can not handle this (like IGV). Nucleotide sequence alignment is a useful comparison technique that detects existing mutations in the DNA sequences.
